High-throughput computing with GROMACS
Note
High-throughput simulations can easily produce a lot of data, so please plan your data management (data flow, storage needs) and analysis pipelines beforehead. Don't hesitate to contact CSC Service Desk if you're unsure about any aspect of your workflow.
GROMACS comes with a built-in multidir functionality, which allows users to run
multiple concurrent simulations within one Slurm allocation. This is an excellent
option for high-throughput use cases, where the aim is to run several similar, but
independent, jobs. Notably, multiple calls of sbatch or srun are not needed,
which decreases the load on the batch queue system. Please consider this option if
you're running high-throughput workflows or enhanced sampling jobs such as replica
exchange or free energy simulations using ensemble-based distance or orientation
restraints.
Another utility of multidir is that it can be used to increase the parallel
efficiency of small systems. By launching multiple trajectories per GPU (or CPU
node), the combined throughput of each independent simulation will increase due
to better resource utilization. This is especially useful for maximizing the
performance of small systems on LUMI-G, as well on Mahti where only full CPU
nodes can be reserved.
Example batch script for Mahti
This example adapts the production part of the lysozyme tutorial
by considering 8 similar copies of the system that have been equilibrated with different velocity
initializations. Inputs corresponding to each copy are named identically md_0_1.tpr
and placed in subdirectories run* as illustrated below by the output of the tree
command.
$ tree
.
├── multidir.sh
├── run1
│ └── md_0_1.tpr
├── run2
│ └── md_0_1.tpr
├── run3
│ └── md_0_1.tpr
├── run4
│ └── md_0_1.tpr
├── run5
│ └── md_0_1.tpr
├── run6
│ └── md_0_1.tpr
├── run7
│ └── md_0_1.tpr
└── run8
└── md_0_1.tpr
#!/bin/bash
#SBATCH --time=00:30:00
#SBATCH --partition=medium
#SBATCH --ntasks-per-node=128
#SBATCH --nodes=1
#SBATCH --account=<project>
# this script runs a 128 core gromacs multidir job (8 simulations, 16 cores per simulation)
module purge
module load gromacs-env
export OMP_NUM_THREADS=1
srun gmx_mpi mdrun -multidir run* -s md_0_1.tpr -dlb yes
By issuing sbatch multidir.sh in the parent directory, all simulations are run concurrently
using one full Mahti node without hyperthreading so that each system is allocated 16 cores.
As the systems were initialized with different velocities, we obtain 8 distinct trajectories
and an improved sampling of the phase space (see RMSD analysis below). This is a great way
to accelerate sampling if your system does not scale to a full Mahti node.
Example batch script for LUMI
Medium-sized and large systems (few 100k–1M+ atoms) are typically able to utilize multiple GPUs on LUMI efficiently. Many smaller use cases run also well on a single GCD (half a GPU), but the smaller the system gets, the poorer it will be able to utilize the full capacity of the accelerator.
The multidir feature can be used to increase the GPU utilization of small systems by
running multiple trajectories per GCD. Below is an example batch script that launches
4 trajectories per GCD in a resource allocation of two GPU nodes (16 GCDs). Each of the
64 .tpr files are organized into separate directories run1 through run64, similar
to the Mahti example above.
#!/bin/bash
#SBATCH --partition=standard-g
#SBATCH --account=<project>
#SBATCH --time=01:00:00
#SBATCH --nodes=2
#SBATCH --gpus-per-node=8
#SBATCH --ntasks-per-node=32
module use /appl/local/csc/modulefiles
module load gromacs/2025.4-gpu
export OMP_NUM_THREADS=1
export MPICH_GPU_SUPPORT_ENABLED=1
export GMX_ENABLE_DIRECT_GPU_COMM=1
export GMX_FORCE_GPU_AWARE_MPI=1
cat << EOF > select_gpu
#!/bin/bash
export ROCR_VISIBLE_DEVICES=\$((SLURM_LOCALID%SLURM_GPUS_PER_NODE))
exec \$*
EOF
chmod +x ./select_gpu
CPU_BIND="mask_cpu:fe000000000000,fe00000000000000"
CPU_BIND="${CPU_BIND},fe0000,fe000000"
CPU_BIND="${CPU_BIND},fe,fe00"
CPU_BIND="${CPU_BIND},fe00000000,fe0000000000"
srun --cpu-bind=${CPU_BIND} ./select_gpu gmx_mpi mdrun -s topol -nb gpu -bonded gpu -pme gpu -update gpu -multidir run*
Note that the number of MPI tasks you request should be a multiple of the number of independent inputs, in this case 1 task per input. Since there are only 56 CPU cores available per LUMI-G node, we use just a single thread per task. For details on the CPU-GPU binding, see the GROMACS application page, as well as LUMI Docs.
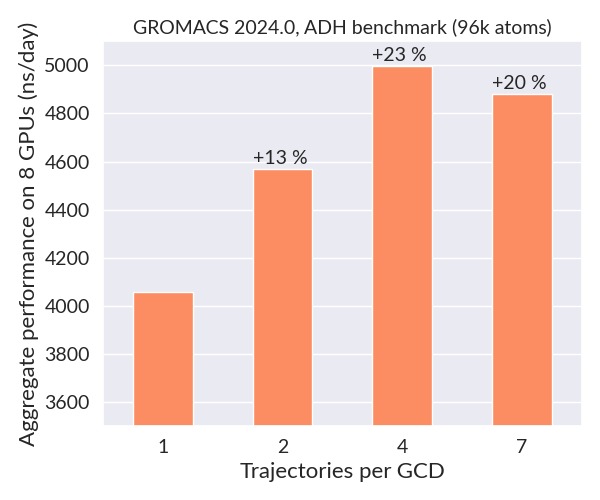
The plot below shows the total combined throughput obtained when running multiple replicas of the 96k atom alcohol dehydrogenase (ADH) benchmark on two LUMI-G nodes (2 fs timestep). When the number of trajectories per GCD is increased from one to four, the overall performance (sum of each independent trajectory) increases by about one microsecond per day due to better GPU utilization. Since each simulation is independent, one could scale this use case to a huge number of nodes for maximal throughput.